Is Medicine Mesmerized by Machine Learning?
BD Horne et al wrote an important paper Exceptional mortality prediction by risk scores from common laboratory tests that apparently garnered little attention, perhaps because it used older technology: standard clinical lab tests and logistic regression. Yet even putting themselves at a significant predictive disadvantage by binning all the continuous lab values into fifths, the authors were able to achieve a validated c-index (AUROC) of 0.87 in predicting death within 30d in a mixed inpatient, outpatient, and emergency department patient population. Their model also predicted 1y and 5y mortality very well, and performed well in a completely independent NHANES cohort1. It also performed very well when evaluated just in outpatients, a group with very low mortality.
1 The authors failed to present a high-resolution validated calibration to demonstrate the absolute predictive accuracy of the model. They also needlessly dealt with sensitivity and specificity.
2 Hemoglobin, red blood count, mean corpuscular hemoglobin, chloride, and BUN were excluded because their information was redundant once all the other predictors were known.
The above model, called by the authors the Intermountain Risk Score, used the following predictors: age, sex, hematocrit, hemoglobin, red cell distribution width, mean corpuscular volume, red blood cell count, platelet count, mean platelet volume, mean corpuscular hemoglobin, mean corpuscular hemoglobin concentration, total white blood count, sodium, potassium, chloride, bicarbonate, calcium, glucose, creatinine, and BUN2. The model is objective, transparent, and needs only one-time and not historical information. It did not need the EHR (other than to get age and sex) but rather used the clinical lab data system. How predicted risks are arrived at is obvious, i.e., a physician can easily see which patient factors were contributing to overall risk of mortality. The predictive factors are measured at obvious times. One can be certain that the model did not use information it shouldn’t such as the use of certain treatments and procedures that may create a kind of circularity with death. It is important to note however that inter-lab variation has created challenges in analyzing lab data from multiple health systems.
Contrast the above under-hyped approach with machine learning (ML). Consider the Avati et al’s paper Improving palliative care with deep learning which was publicized here. The Avati paper addresses an important area and is well motivated. Palliative care (e.g., hospice) is often sought at the wrong time and relies on individual physician referrals. An automatic screening method may yield a list of candidate patients near end of life who should be evaluated by a physician for the possibility of recommending palliative rather than curative care. A method designed to screen for such patients needs to be able to estimate either mortality risk or life expectancy accurately.
Avati et al’s analysis used a year’s worth of prior data on each patient and was based on 13,654 candidate features from the EHR. As with any retrospective study not based on an inception cohort with a well-defined “time zero”, it is tricky to define a time zero and somewhat easy to have survival bias and other sampling biases sneak into the analysis. The ML algorithm, in order to use a binary outcome, required division of patients into “positive” and “negative” cases, something not required by regression models for time until an event3. “Positive” cases must have at least 12 months of previous data in the health system, weeding out patients who died quickly. “Negative” cases must have been alive for at least 12 months from the prediction date. It is also not clear how variable censoring times were handled. In standard statistical model, patients entering the system just before the data analysis have short follow-up and are right-censored early, but still contribute some information.
3 There exist neural network algorithms for censored time-to-event data.
Avati et al used deep learning on the 13,654 features to achieve a validated c-index of 0.93. To the authors’ credit, they constructed an unbiased calibration curve, although it used binning and is very low resolution. Like many applications of ML where few statistical principles are incorporated into the algorithm, the result is a failure to make accurate predictions on the absolute risk scale. The calibration curve is far from the line of identity as shown below.
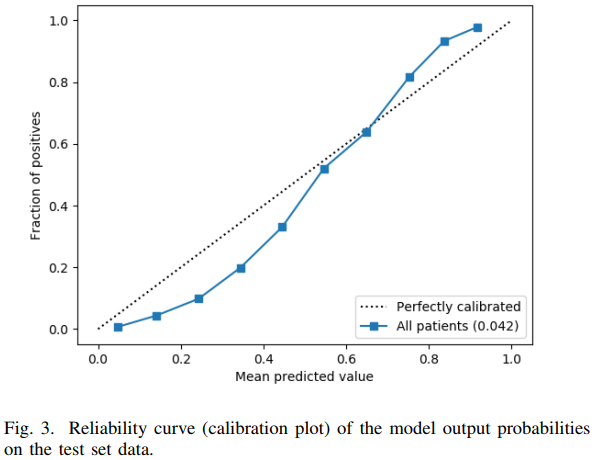
The authors interpreted the above figure as “reasonably calibrated.” It is not. For example, a patient with a predicted probability of 0.2 had an actual risk < 0.1. The gain in c-index from ML over simpler approaches has been more than offset by worse calibration accuracy than the other approaches achieved.
Importantly, some of the hype over ML comes from journals and professional societies and not so much from the researchers themselves. That is the case for the Avati et al deep learning algorithm, which is not actually being used in production mode at Stanford. A much better calibrated and somewhat more statistically-based algorithm is currently being used.
Like many ML algorithms, the focus is on development of “classifiers”. As detailed here, classifiers are far from optimal in medical decision support where decisions are not to be made in a paper but only once utilities/costs are known. Utilities and costs only become known during the physician/patient interaction. Unlike statistical models which directly estimate risk or life expectancy, the majority of ML algorithms start by using classification, then if a probability is needed they try to convert the patterns into a probability (this is sometimes called a “probability machine”). As judged by Avati et al’s calibration plot, this conversion may not be reliable.
Avati et al, besides showing us what is needed, and consistent with forward prediction (the calibration plot) also reported a number of problematic measures. As detailed here, the use of improper probability accuracy scoring rules is very common in the ML world, because of the hope that one can actually make a decision (classification) using the data without needing to incorporate costs of incorrect decisions (utilities). Improper accuracy scores have a number of problems, such as
- reversing information flow, i.e., conditioning on outcomes and examining tendencies of inputs
- inviting dichotomization of inputs
- being optimized by choosing the wrong features and giving them the wrong weights
Proportion classified correctly, sensitivity, specificity, precision, and recall are all improper accuracy scoring rules and should not play a role in a forward prediction mode when risk or life expectancy estimation are the real goals. A poker player wins consistently because she is able to estimate the probability she will ultimately win with her current hand, not because she recalls how often she’s had such a hand when she won.
One additional point: the ML deep learning algorithm is a black box, not provided by Avati et al, and apparently not usable by others. And the algorithm is so complex (especially with its extreme usage of procedure codes) that one can’t be certain that it didn’t use proxies for private insurance coverage, raising a possible ethics flag. In general, any bias that exists in the health system may be represented in the EHR, and an EHR-wide ML algorithm has a chance of perpetuating that bias in future medical decisions. On a separate note, I would favor using comprehensive comorbidity indexes and severity of disease measures over doing a free-range exploration of ICD-9 codes.
It may also be useful to contrast the ML approach with another carefully designed traditional and transparent statistical approach used in the HELP study of JM Teno, FE Harrell, et al. A validated parametric survival model was turned into an easy-to-use nomogram for obtaining a variety of predictions on older hospitalized adults:
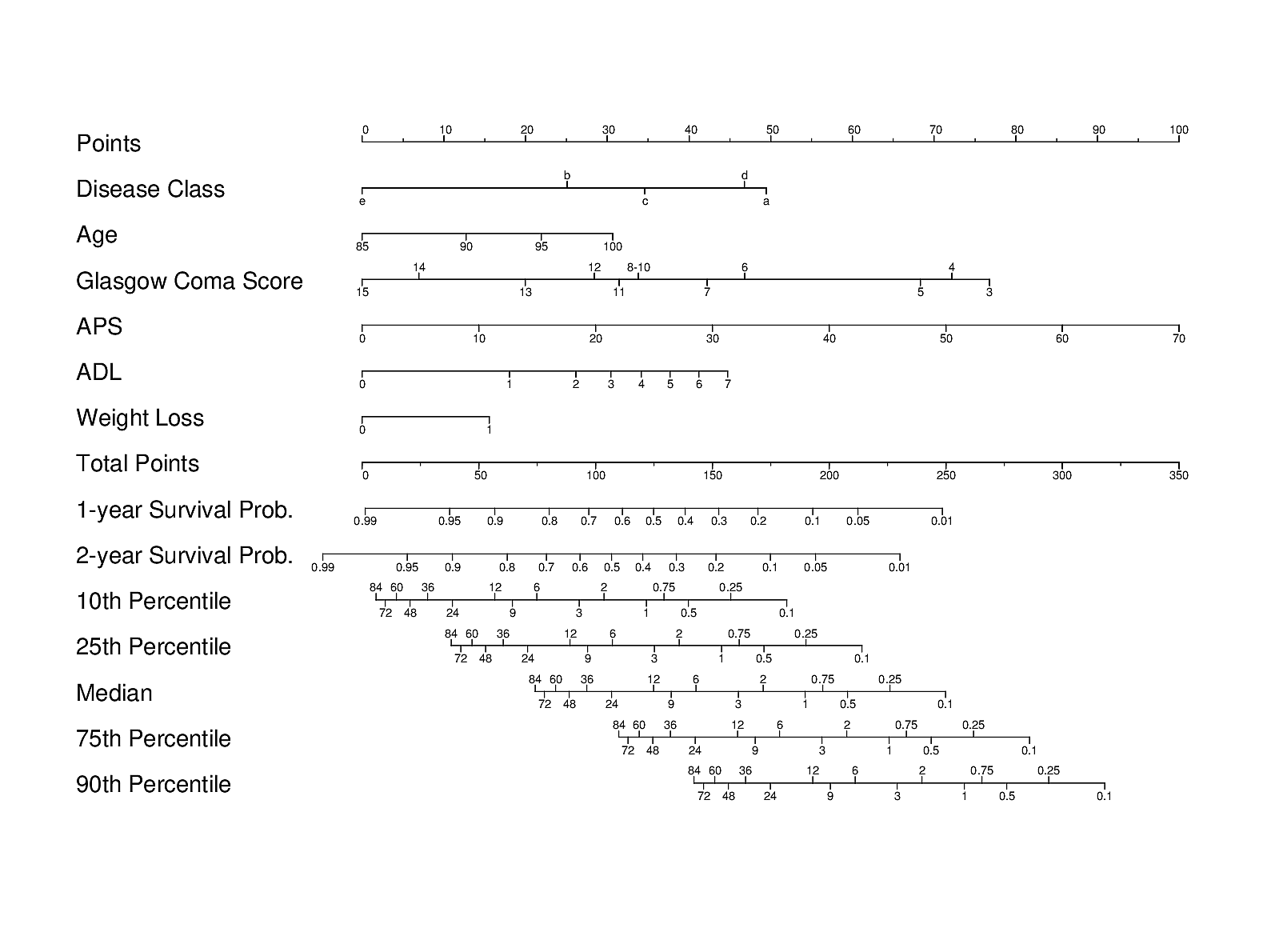
Importantly, patients’ actual preferences for care were also studied in HELP. A different validated prognostic tool for end-of-life decision making, derived primarily from ICU patients, is the SUPPORT prognostic model.
In the rush to use ML and large EHR databases to accelerate learning from data, researchers often forget about the advantages of statistical models and of using more compact, cleaner, and better defined data. They also sometimes forget how to measure absolute predictive accuracy, or that utilities must be incorporated to make optimum decisions. Utilities are applied to predicted risks; classifiers are at odds with optimum decision making and with incorporating utilities at the appropriate time, which is usually at the last minute just before the medical decision is made and not when a classifier is being built.
References: Guidelines for Reporting Predictive Models
Other Relevant Articles
- Rage Against the Machine Learning by Maarten van Smeden
- Poor Reporting, Weak Methods, High Risk of Bias, Full of Spin by Gary Collins
- Big Data and Machine Learning in Health Care
- UK Parliament AI Report
- Statistical and Machine Learning Forecasting Methods: Concerns and Ways Forward by S Markridakis, E Spiliotis, V Assimakopoulos
- Excellent discussion of overfitting, measuring accuracy, and lack of rigor in published machine learning studies in financial time series forecasting. Simple statistical methods outperformed complex machine learning algorithms. Previous researchers refused to share data.
- Big Data and Predictive Analytics: Recalibrating Expectations by ND Shah, EW Steyerberg, DM Kent
- What are radiological deep learning models actually learning? by John Zech
- Biases in electronic health record data due to processes within the healthcare system: retrospective observational study by Denis Agniel et al.
- Data about timing of medical test ordering was more predictive of survival than the actual test results
- Special issue on probability estimation and machine learning of Biometrical Journal, including discussion articles comparing ML and SM
- Questions for artificial intelligence in health care by Maddox et al
- High-performance medicine: the convergence of human and artificial intelligence by Eric Topol
- The exaggerated promise of so-called unbiased data mining
- A systematic review shows no performance benefit of machine learning over logistic regression for clinical prediction models by Evangelia Christodoulou et al.
- Criminal machine learning by Carl Bergstrom and Jevin West
- Machine learning, practically speaking by Vivien Marx
- Stop explaining black box machine learning models for high stakes decisions and use interpretable models instead by Cynthia Rudin
- Why are we using black box models in AI when we don’t need to? A lesson from an explainable AI competition by Cynthia Rudin and Joanna Radin
- Artificial intelligence could revolutionize medical care. But don’t trust it to read your x-ray just yet by Jennifer Couzin-Frankel
- Subject matter knowledge in the age of big data and machine learning by Ben Goldstein et al.
- Three pitfalls to avoid in machine learning by Patrick Riley
- Association between surgical skin markings in dermoscopic dmages and diagnostic performance of a deep learning convolutional neural network for melanoma recognition by Julia Winkler et al
- The “inconvenient truth” about AI in healthcare by T Panch, H Mattie, L Celi
- Deep learning: A critical appraisal by Gary Marcus
- Comments on Rebooting AI by Eric Topol
- AI equal with human experts in medical diagnosis, study finds by Nicola Davis
- Why deep-learning AIs are so easy to fool by Douglas Heaven
- Hidden stratification causes clinically meaningful failures in machine learning for medical imaging by Luke Oakden-Rayner et al.
- Machine learning for prediction in electronic health data by Sherri Rose
- Your evidence? Machine learning algorithms for medical diagnosis and prediction by Bert Heinrichs and Simon Eickhoff
- Andreessen-Horowitz on AI startups
- Fair warning by Adeba Birhane
- Logistic regression was as good as machine learning for predicting major chronic diseases by Nusinovici et al
- AI for medicine is overhyped right now by Dan Elton
- Is AI leading to a reproducibility crisis in science? by Philip Ball